import urllib.request
# Download the image and save it as a local file
with urllib.request.urlopen("https://psiclone.tfda.jp/asset/images/toymodel.png") as web_file:
with open("toymodel.png", "wb") as local_file:
local_file.write(web_file.read())Analysis of Image Data
Loading the Data
Let us analyze the following grayscale image toymodel.png.
Load the image as a NumPy array using cv2 (OpenCV).
import cv2
import numpy as np
image_file_path = "./toymodel.png"
levels = np.array(cv2.imread(image_file_path, cv2.IMREAD_GRAYSCALE), dtype=np.float64)Load levels into a psiclone data structure.
import psiclone.data
img = psiclone.data.GrayscaleImage(levels)Draw the loaded img to verify that it has been read correctly.
import matplotlib.pyplot as plt
import psiclone.visualization as vis
vis.draw_geometry(
img,
bitmap=True,
)
plt.colorbar()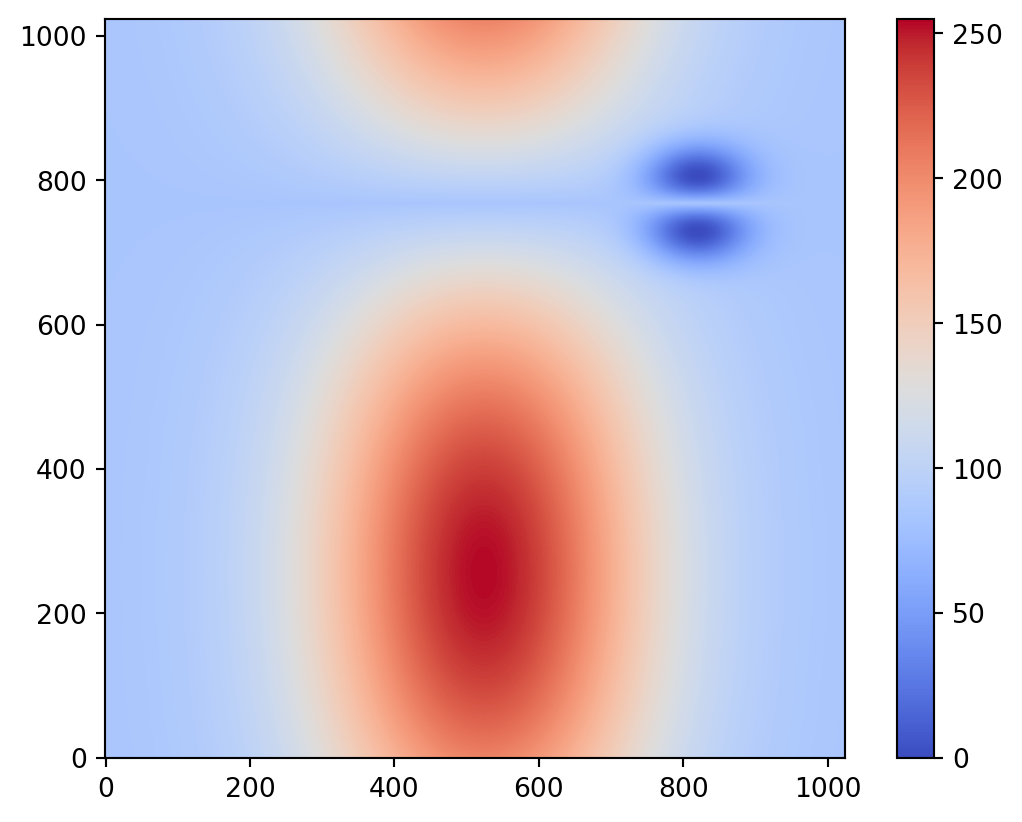
Converting the Input Data to a COT Representation
Analyze the loaded img and compute its partially Cyclically Ordered rooted Tree (COT) representation.
This makes use of the psiclone.psi2tree.Psi2tree class.
Depending on the input data, the computation may take a long time. Therefore, you may need to set an appropriate noise-removal threshold (threshold).
If the computation takes too long (or if the resulting COT representation is unnecessarily detailed), raise the threshold appropriately, e.g., to 1.0 or 10.
from psiclone.psi2tree import Psi2tree
psi2tree = Psi2tree(img, threshold=0)
psi2tree.compute()After computing the COT representation, obtain the result using the get_tree method.
The tree structure data returned by get_tree can then be converted into a human-readable format using the convert method.
from psiclone.cot import to_str
psi2tree.get_tree().convert(to_str)'a0(a-(b--{s-,s-}).a+(s+))'By passing one of the following functions to the convert method, the COT representation can be transformed into various output formats:
psiclone.cot.to_str- Converts to a simple ASCII format without Greek letters
psiclone.cot.to_short_str- Similar to
psiclone.cot.to_str, but omits redundant symbols without losing information
- Similar to
psiclone.cot.to_tex- Converts to a TeX format with correct subscripts and Greek-letter labels
psiclone.cot.to_short_tex- Similar to
psiclone.cot.to_tex, but omits redundant symbols without losing information
- Similar to
You can also customize the recursive transformation by changing the function passed to convert.
from IPython.display import Math, display
from psiclone.cot import to_short_str, to_short_tex
cot_tree = psi2tree.get_tree()
print(cot_tree.convert(to_short_str))
print(cot_tree.convert(to_short_tex))
display(Math(cot_tree.convert(to_short_tex)))a0(a-(b--).a+)
a_\emptyset(a_-(b_{--})\cdot a_+)Visualization of the Results
Let us overlay the input data (levels), contour plot, Reeb graph, and COT representation.
# 0. Set image resolution
plt.rcParams["figure.figsize"] = [5, 5]
plt.rcParams["figure.dpi"] = 100
# 1. Display the COT representation at the top of the image
plt.title(f"${psi2tree.get_tree().convert(to_short_tex)}$")
# 2. Draw the input data (levels) and contour plot
y = list(range(levels.shape[0]))
x = list(range(levels.shape[1]))
plt.imshow(levels, cmap="coolwarm", origin="lower", extent=[np.amin(x), np.amax(x), np.amin(y), np.amax(y)])
plt.contour(x, y, levels, colors="k")
# 3. Overlay the Reeb graph and COT representation
vis.draw_graph_markers(psi2tree, lw=2, edge_color="y-", node_color="g*")
vis.draw_COT_labels(psi2tree, fontsize=16)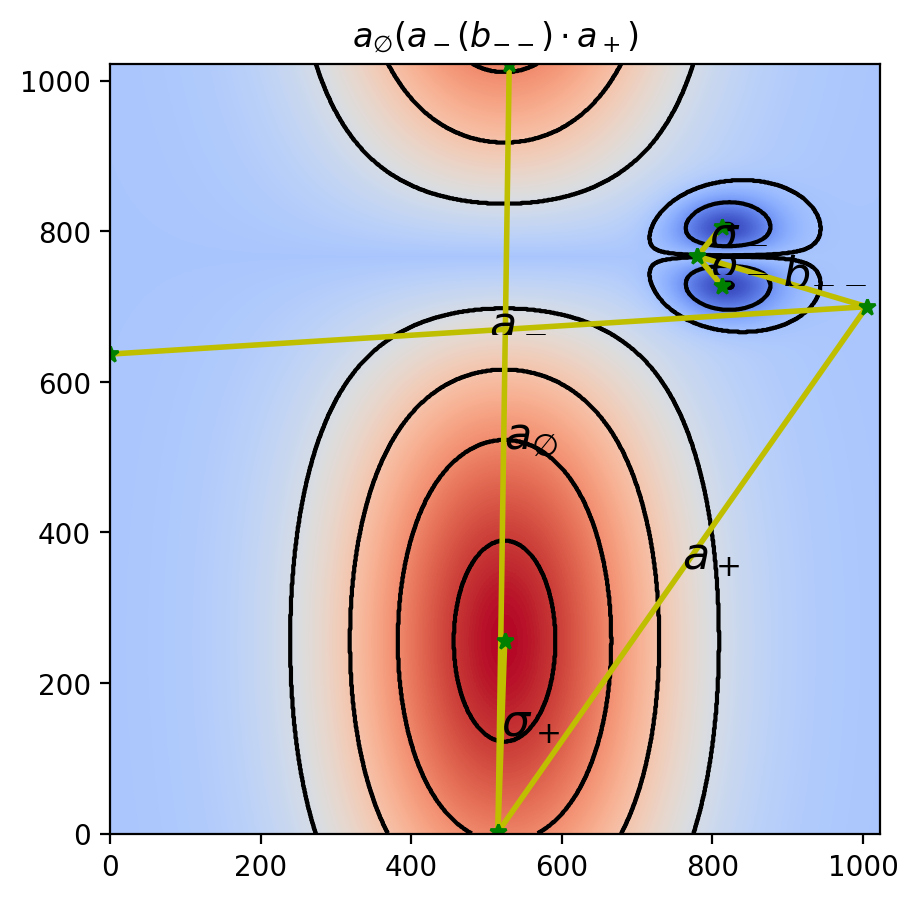
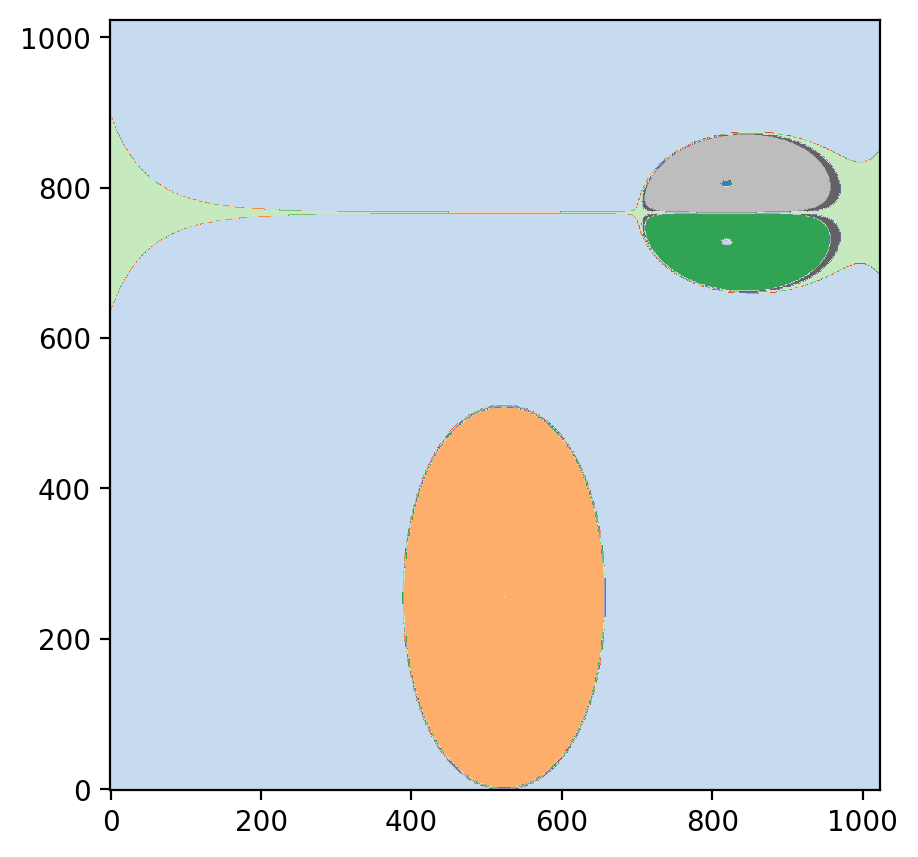
Visualization of the Partition
Mathematically, a Reeb graph is defined as a quotient space under an equivalence relation.
The algorithm used in psiclone also produces a partition of the space based on such an equivalence relation.
To visualize the partition, the psiclone.visualization.draw_partition function is convenient.
By examining this, you can get an intuitive understanding of how psiclone captures streamline topology through the Reeb graph.
plt.rcParams["figure.figsize"] = [5, 5]
plt.rcParams["figure.dpi"] = 100
vis.draw_partition(psi2tree, bitmap=True, discrete=True, cmap="tab20c")